  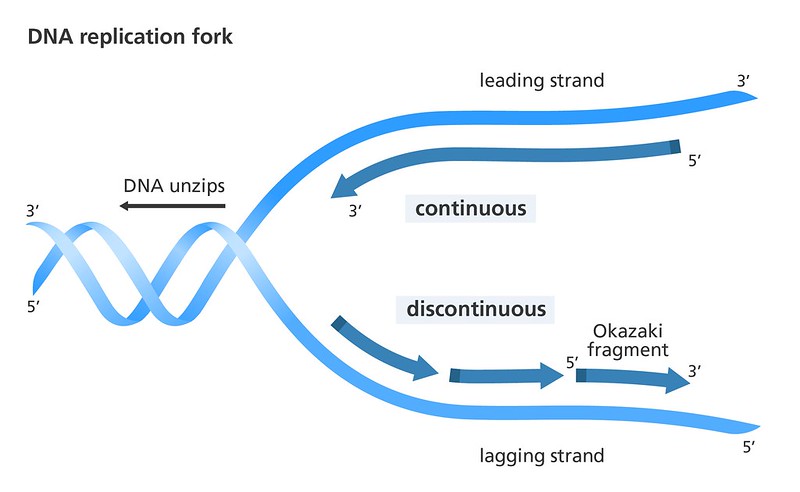 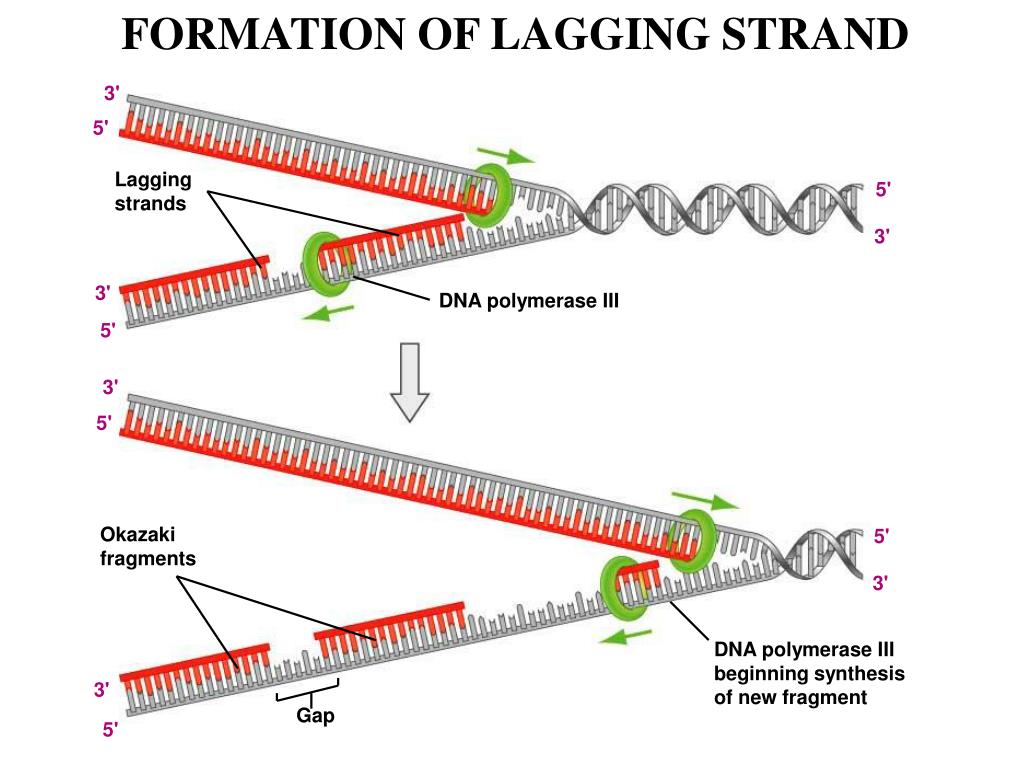
Polymerases confine the mechanisms that can be used by the cell for DNA duplication. The antiparallel structure of double-helical DNA and the 3′ end extension specificity of all DNA Replication of cellular chromosomal DNA is initiated by the multienzyme replisome machinery, which unwinds the DNA helix toĬreate a replication fork. That can shift from high efficiency to high fidelity. The eukaryotic maturation mechanism involves many enzymes, possibly three pathways, and regulation The eukaryotic fragments are much shorter, with lengths determined by nucleosome periodicity. 
Although the prokaryotic fragments are ∼1200 nucleotides long, Of the primer into a flap, flap removal, and then ligation. The lagging-strand fragments are initiated by RNA primers, which are removed by a joining mechanism involving strand displacement Identified proteins and multiple pathways responsible for maturation of the lagging strand. Genetic analyses and reconstitution experiments The lagging strand needs to be processed to form a functional DNA segment. The leading strand is elongatedĬontinuously in the direction of fork opening, whereas the lagging strand is made discontinuously in the opposite direction. Okazaki fragments are a part of the lagging strand, and lagging strands give space to Okazaki fragments which is responsible for the continuous growth of Okazaki fragments.Cellular DNA replication requires efficient copying of the double-stranded chromosomal DNA.While lagging strands continue formation with the short RNA sequences and maintain the development of Okazaki fragments. Okazaki fragments are formed by the action undertaken by the strand.Okazaki fragments are 100 to 200 nucleotides long, but the average size of lagging strands is 3000 nucleotides long.Okazaki fragments are comparatively shorter in size, but lagging strands, when compared, are much longer in size.It is responsible for the continuous growth of Okazaki fragments. Okazaki Fragments are small sections of DNA that are dependent on lagging strands to be produced, whereas lagging strands are nothing but replicated DNA strands.Main Differences Between Okazaki Fragments and Lagging Strand Without the work of lagging strands, the process of making Okazaki fragments will get disrupted, and ultimately the proper DNA formation will be compromised. DNA ligase is another requirement in the joining of Okazaki fragments. The formation of Okazaki fragments requires a new RNA primer every time. When lagging strands start to synthesize, they produce Okazaki fragments. The formation of lagging strands is a slow process. It is capable of opening up in the 5′ to 3′ direction and grows in the 3′ to 5′ direction. The growth of lagging strands takes place when it discontinuously constructs Okazaki Fragments. Japanese biologists Tsuneko Okazaki discovered them in the 1960s with the help of Reiji and other colleagues. These fragments are named after their founder. These are short, roughly we can measure their average size to be 150 to 200 nucleotides. Okazaki fragments are cycles of DNA nucleotides. Lagging strands give space to Okazaki fragments. Found in It is a part of the lagging strand. Its average size is 3000 nucleotides long. Size These are 100 to 200 nucleotides long. Lagging strands continue formation with the short RNA sequences and maintain the development of Okazaki fragments. Formation It is formed by the action undertaken by the strand. Length It is comparatively shorter in size. Comparison Table Parameters of Comparison Okazaki Fragments Lagging Strand Definition These are small sections of DNA that are produced by the uninterrupted process of lagging strands. These are large in size, and it is the other half of the leading strand which grows without a gap, unlike lagging strands. The formation of Okazaki fragments is not possible unless lagging strands perform their duties. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
